apply() function
How it works
Example Data
This is the value of matrix a
## [,1] [,2] [,3]
## A 1 6 11
## B 2 7 12
## C 3 8 13
## D 4 9 14
## E 5 10 15Operation with built-in functions
The mean of a’s rows
apply(a,1, mean)## A B C D E
## 6 7 8 9 10The stddev of a’s rows
apply(a,1, sd)## A B C D E
## 5 5 5 5 5For common operations such as sum, mean, there are dedicated, vectorized versions which are faster.
all(apply(a, 1, mean) == rowMeans(a))## [1] TRUEall(apply(a, 2, mean) == colMeans(a))## [1] TRUEall(apply(a, 1, sum) == rowSums(a))## [1] TRUEall(apply(a, 2, sum) == colSums(a))## [1] TRUE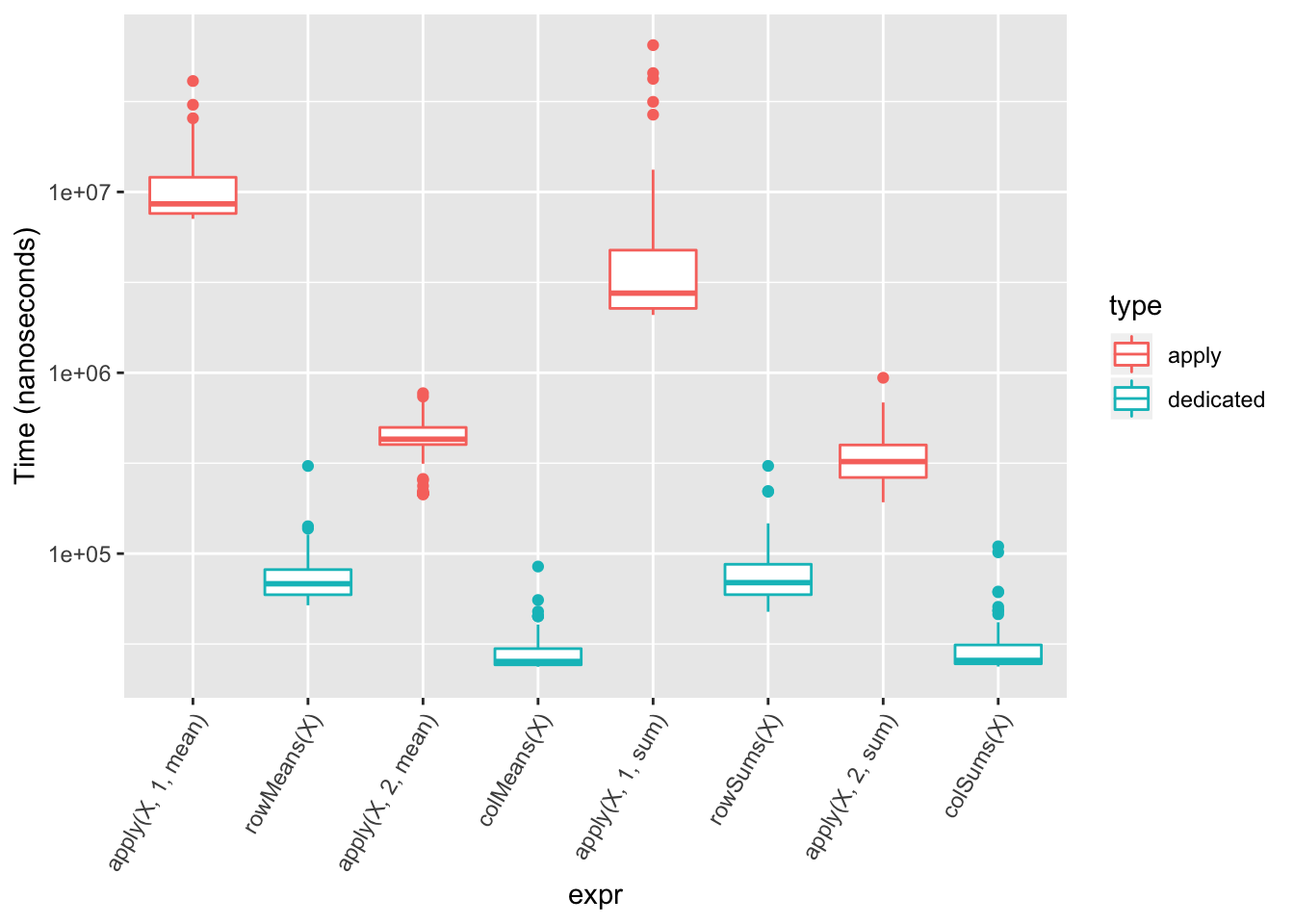
Operation with custom-defined function: z-score conversion
z0 function is defined to normalize a vector against its own mean and stddev
z0= function(x) {return((x-mean(x)) / sd(x))}z1 function is defined to normalize a vector against another mean and stddev
z1= function(x) {return((x-xm)/xs )}Applying z0() along the rows of a gives
apply(a,1,z0)## A B C D E
## [1,] -1 -1 -1 -1 -1
## [2,] 0 0 0 0 0
## [3,] 1 1 1 1 1apply(t(a), 2, z0)## A B C D E
## [1,] -1 -1 -1 -1 -1
## [2,] 0 0 0 0 0
## [3,] 1 1 1 1 1Applying z0() along the columns of a gives
apply(a,2,z0)## [,1] [,2] [,3]
## A -1.2649111 -1.2649111 -1.2649111
## B -0.6324555 -0.6324555 -0.6324555
## C 0.0000000 0.0000000 0.0000000
## D 0.6324555 0.6324555 0.6324555
## E 1.2649111 1.2649111 1.2649111The result of apply() along the row (axis=1) of the matrix is transposed since each operation return a column vector, and these vectors are cbind to give the final matrix.
Applying z1() with precalculated mean and sd gives
xm = c(6,7,8,9,10)
xs = c(5,5,5,5,5)
apply(a,2,z1)## [,1] [,2] [,3]
## A -1 0 1
## B -1 0 1
## C -1 0 1
## D -1 0 1
## E -1 0 1apply(t(a),2,z1)## Warning in x - xm: longer object length is not a multiple of shorter object
## length
## Warning in x - xm: longer object length is not a multiple of shorter object
## length
## Warning in x - xm: longer object length is not a multiple of shorter object
## length
## Warning in x - xm: longer object length is not a multiple of shorter object
## length
## Warning in x - xm: longer object length is not a multiple of shorter object
## length## A B C D E
## [1,] -1.0 -0.8 -0.6 -0.4 -0.2
## [2,] -0.2 0.0 0.2 0.4 0.6
## [3,] 0.6 0.8 1.0 1.2 1.4
## [4,] -1.6 -1.4 -1.2 -1.0 -0.8
## [5,] -0.8 -0.6 -0.4 -0.2 0.0Performance
Do the built-in apply functions perform better than a for loop? Not quite. The example with sapply below showed that it’s even a bit behind a regular for loop. That means apply functions only provide a short-hand for the serial looping. You’d still need parallelization to improve performance.
complicatedOp <- function(x) {
return(log10(x^x^x) + x^2 / x*4.128 + sqrt(x)*(x^(0.31234)) )
}apply.for = function(anArray) {
output = rep(0,length(anArray))
for (i in 1:length(anArray)) {
output[i] = complicatedOp(anArray[i])
}
}
apply.builtin = function(anArray) {
output = sapply(anArray, FUN = complicatedOp)
}
apply.vectorized = function(anArray) {
output = vapply(anArray, FUN.VALUE = 0., FUN = complicatedOp)
}sizes = c(1000, 5000,10000)
a1= runif(sizes[1])
a2= runif(sizes[2])
a3= runif(sizes[3])
bm = microbenchmark::microbenchmark(apply.for(a1),
apply.for(a2),
apply.for(a3),
apply.builtin(a1),
apply.builtin(a2),
apply.builtin(a3),
apply.vectorized(a1),
apply.vectorized(a2),
apply.vectorized(a3))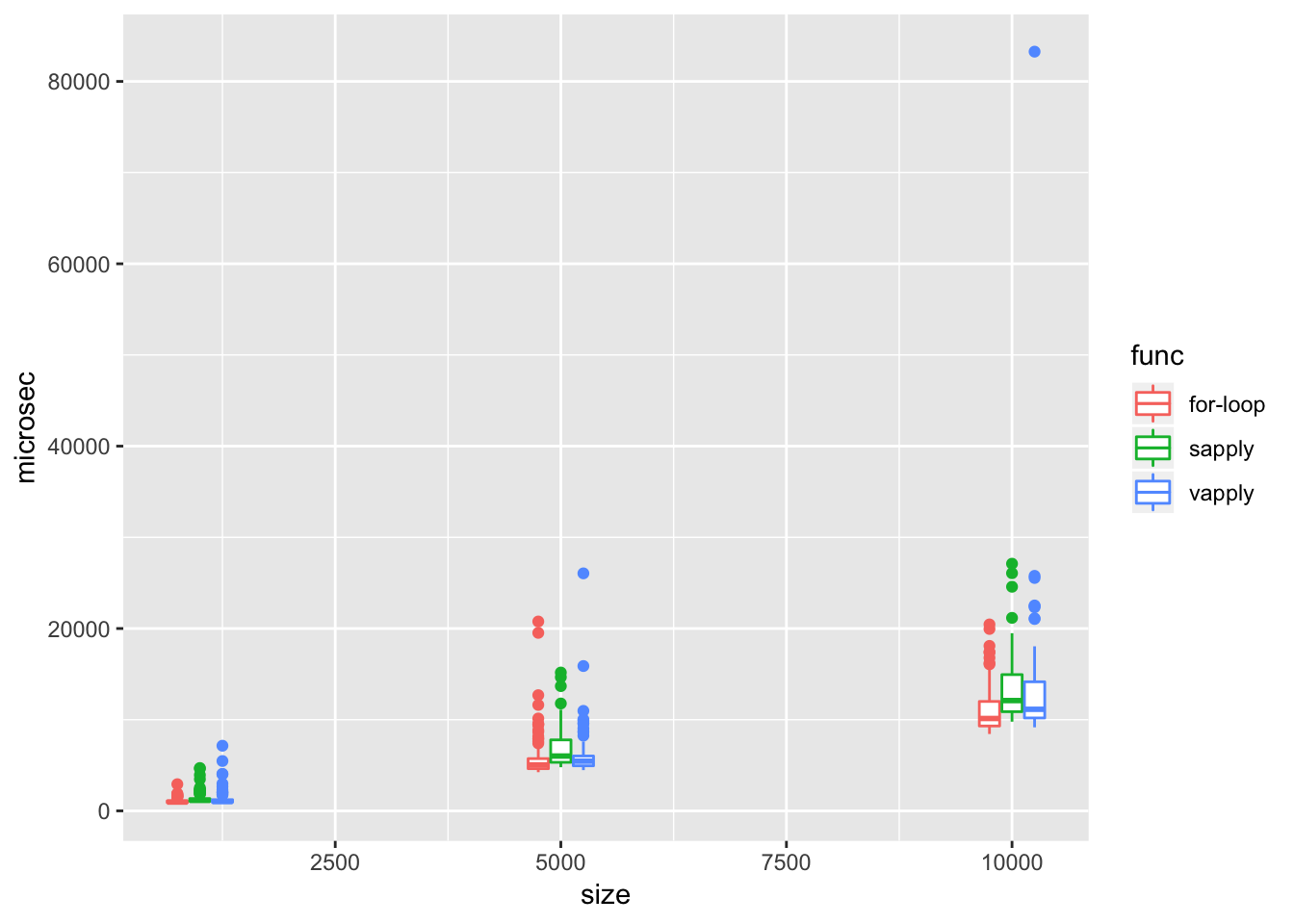
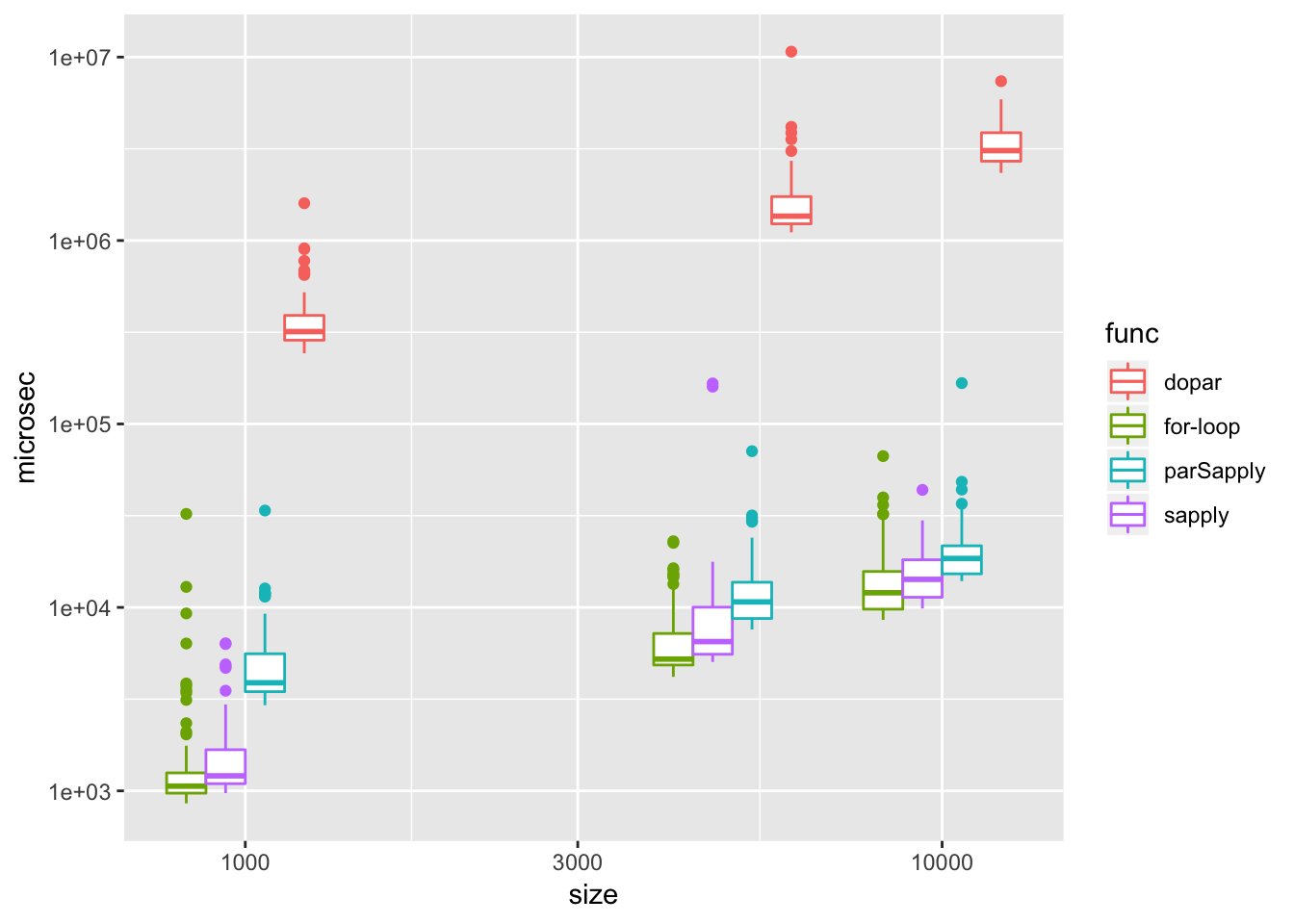
library(parallel)
cl = makeCluster(4)
apply.par1 = function(anArray) {
output = parSapply(cl,anArray, FUN = complicatedOp)
}
library(doParallel)
apply.par2 = function(anArray) {
registerDoParallel(cores=6)
output = rep(0,length(anArray))
foreach (i=1:length(anArray)) %dopar% {
output[i] = complicatedOp(anArray[i])
}
}bm2 = microbenchmark::microbenchmark(apply.for(a1),
apply.for(a2),
apply.for(a3),
apply.builtin(a1),
apply.builtin(a2),
apply.builtin(a3),
apply.par1(a1),
apply.par1(a2),
apply.par1(a3),
apply.par2(a1),
apply.par2(a2),
apply.par2(a3))